Carlos Perea Resa Lab. 409 Ext. 4463
Dynamic and Recycling of the transcriptional machinery across mitosis.
One of the most exciting aspects of our biology is the fact that all our cells share the same genetic information (handbook). The differential reading of this info is key to generate the diversity of cell types during the developmental program of a complex organism. In this frame, the transcription of DNA into RNA constitutes the first stage of that reading and is essential to establish differential gene expression patterns defining the distinct cell lineages. In addition to transcription, the process of mitosis is fundamental to obtain the precise number of cells to build a whole organism while the establishment of most of the cell lineages required the passage of cells through mitosis. Interestingly, transcription is widely silenced during mitosis due the release of RNA polymerases and transcription factors from the condensing chromatin. Once mitosis ends, transcription restart immediately to guarantee the viability and identity of daughter cells. The mechanisms behind transcription restart following mitosis are poorly understood. How daughter cells remember the transcriptional program of the mother or/and how those programs rewire during development are mostly unknown. In our laboratory we aim to answer these questions using the culture of mammalian cells as a model. One of our goals is to understand the dynamic and recycling of the RNA polymerase 2, the enzyme transcribing all protein coding genes in eukaryotes, and the associated complex named Cohesin. In parallel, we investigate the potential implication of these processes in the etiology of developmental diseases as cohesinopathies. By combining the use of cell culture, genomic and proteomic approaches, CRISPR technology and live cell microscopy, we aim to understand how the transcriptional machinery works across the challenge of mitosis with the hope to understand how gene expression and cell division coordinates during development.
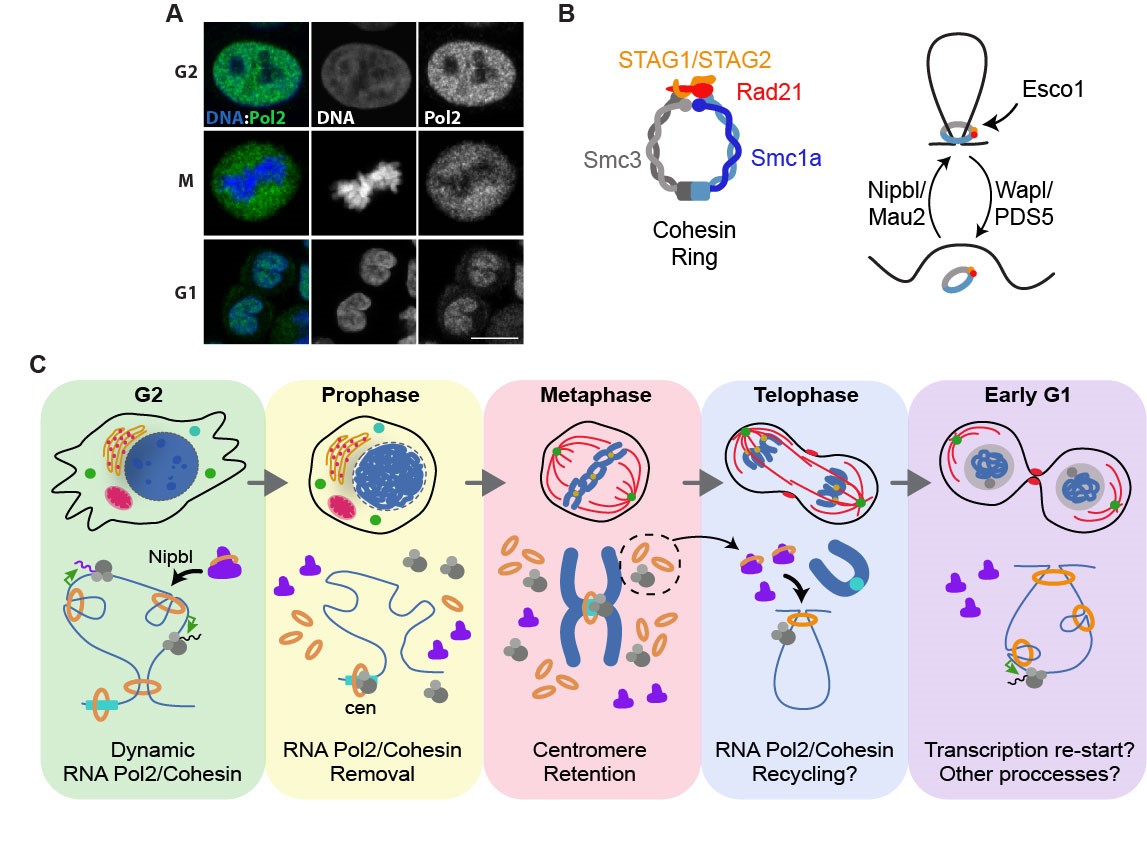
Figure 1. Dynamic and Recycling of the transcriptional machinery across mitosis. A Subcellular localization of RNA Pol2 in G2, M and G1 stages by immunofluorescence. B Schematic representation of the cohesin complex (left) and the regulators of its interaction with chromatin (right). C Model representing the dynamic and potential recycling of RNA Pol2 (grey) and cohesin (orange) across mitosis.
Twitter: @PereaResa

| Last name | Name | Laboratory | Ext.* | Professional category | |
|---|---|---|---|---|---|
| Contreras Pérez | Aida | 322 | 4611 | aida.contreras(at)cbm.csic.es | M3 Predoc.formación |
| García Piñero | María | 409 | 4611 | Estudiante TFM | |
| Perea Resa | Carlos | 409 | 4611 | carlos.perea(at)cbm.csic.es | Doctor FC1 |
| Sayago Pachón | Estrella Isabel | 409 | 4611 | Becario JAE Intro |
1. Perea-Resa C*, Wattendorf L, Marzouk S, and Blower MD*. Trends in Cell Biology, 2021 doi: 10.1016/j.tcb.2021.03.005. *CO-CORRESPONDING AUTHORS
2. Ardehali MB, Damie M, Perea-Resa C, Blower MD and Kingston RE. Journal of Biological Chemistry, 2020. doi: 10.1074/jbc.RA120.015877
3. Hekman RM, Hume AJ, Raghuveera KG, et al., Actionable Cytopathogenic Host Responses of Human Alveolar Type 2 Cells to SARS-CoV-2. Molecular Cell, 2020. doi:10.1016/j.molcel.2020.11.028
4. Sharp JA, Perea-Resa C, Wang W, Blower MD. Cell division requires RNA eviction from condensing chromosomes. Journal of Cell Biology, 2020. doi: 10.1083/jcb.201910148
5. Perea-Resa C, Bury L, Cheeseman IM, Blower MD. Cohesin Removal Reprograms Gene Expression upon Mitotic Entry. Molecular Cell, 2020. doi: 10.1016/j.molcel.2020.01.023
6. Esteve-Bruna D, Carrasco-López C, Blanco-Touriñán N, et al., Prefoldins contribute to maintaining the levels of the spliceosome LSM2-8 complex through Hsp90 in Arabidopsis. Nucleic Acids Research, 2020. doi: 10.1093/nar/gkaa354
7. Perea-Resa C, Blower MD. Centromere Biology: Transcription Goes on Stage. Molecular and Cell Biology, 2018. Review. doi: 10.1128/MCB.00263-18
8. Perea-Resa C, Blower MD. Satellite Transcripts Locally Promote Centromere Formation. Developmental Cell, 2017. doi: 10.1016/j.devcel.2017.07.017
9. Perea-Resa C, Rodríguez-Milla MA, Iniesto E, Rubio V, Salinas J. Prefoldins negatively regulate cold acclimation in Arabidopsis thaliana by promoting nuclear proteasome-mediated HY5 degradation. Molecular Plant, 2017. doi: 10.1016/j.molp.2017.03.012
10. Carrasco-López C, Hernández-Verdeja T, Perea-Resa C, Abia D, Catalá R, Salinas J. Environmental-dependent regulation of spliceosome activity by the LSM2-8 complex in Arabidopsis. Nucleic Acids Research, 2017. doi: 10.1093/nar/gkx375
11. Perea-Resa C, Carrasco-López C, Catalá R, Turečkova V, Novak O, Zhang W, Sieburth L, Jiménez-Gómez JM, Salinas J. The LSM1-7 complex differentially regulates Arabidopsis thaliana tolerance to abiotic stress conditions by promoting selective mRNA decapping. The Plant Cell, 2016. doi: 10.1105/tpc.15.00867
12. Perea-Resa C, Hernández-Verdeja T, López-Cobollo R, del Mar Castellano M, Salinas J. LSM proteins provide accurate splicing and decay of selected transcripts to ensure normal Arabidopsis development. The Plant Cell, 2012. doi: 10.1105/tpc.112.103697




















