Genomic approaches to identify genes implicated across major psychiatric disorders
Research summary:
The identification of genes in psychiatric diseases is our front-line research. Family studies have established a strong genetic contribution for most psychiatric diseases, but the specific genes involved still remain largely unknown.
Genome-wide approaches indicate that psychiatric disorders are likely to be caused by a combination of common variants, each with a small effect, and multiple rare variants of higher penetrance. In our group we adopt several genomic approaches and a combination of them to identify susceptibility genes implicated in autism spectrum disorder (ASD), bipolar disorder (BD), and schizophrenia (SCZ):
- Linkage studies: This methodology aims to map disease-causing loci by analysing polymorphic DNA markers that co-segregate with the phenotype of interest in multiplex families.
- Genome-wide association studies (GWAS): this approach is used to identify Single Nucleotide Variants (SNPs) (common DNA variation) associated with the disorder.
- Copy number variant (CNV) analysis: small deletions or duplications of genetic material (>50bp) particularly those spanning genes may represent etiologic causes for disease.
- Next Generation Sequencing (NGS): whole exome sequencing (WES) and whole genome sequencing (WGS) are able to identify rare single nucleotide variants (SNVs) and de novo (DN) (rare DNA variation) that may impair the normal function of proteins, and increase risk of disease.
Our expertise in high-throughput data analyses identified several genes in psychiatric disorders: i) The X-linked DRP2 gene is now included in a panel for genetic testing of autism with intellectual disability (ID); ii) we found the YHWAZ gene implicated in autism; iii) we identified IRS4 gene disrupted in bipolar disorder; iv) We showed the involvement of LRP1 gene in schizophrenia via common variants and autism via rare variants; v) we showed that higher burden of truncating variants (alleles that disrupt proteins) are associated with severe symptomatology in autism and bipolar disorder, and may modulate cognitive function; vi) We demonstrated that linkage studies are powerful approaches when combined with whole-exome sequencing (WES) in complex disorders (polygenic inheritance).
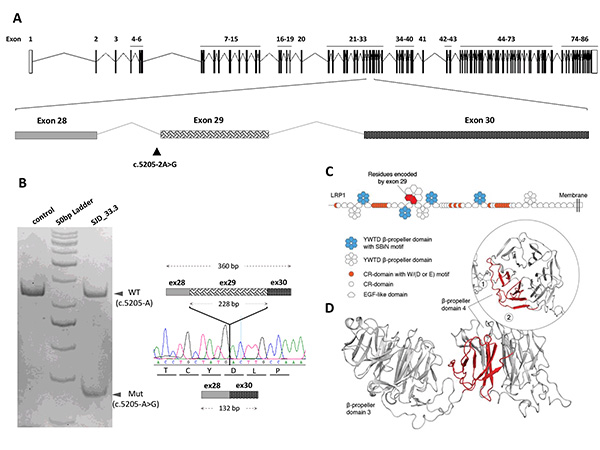
Figure 1. Whole-exome sequencing identified a de novo variant with splicing effect on LRP1 in an autistic patient. (A) The mutation site is indicated by a triangle (c.5205–2A>G) between exons 28 to 30. (B) Polymerase chain reaction (PCR) on the mutated transcript (Mut) generated a smaller fragment (132 bp) lacking exon 29 (76 amino acids), which generates an in-frame transcript. (C) LRP1 domains with the β-propeller domains as hexagons. The region encoded by exon 29 in β-propeller 4 is in red. (D) The skipping of exon 29 led to the removal of the first 2 blades of 6 from β-propeller 4. (Torrico,…Toma (2019) Journal of Psychiatry and Neuroscience).
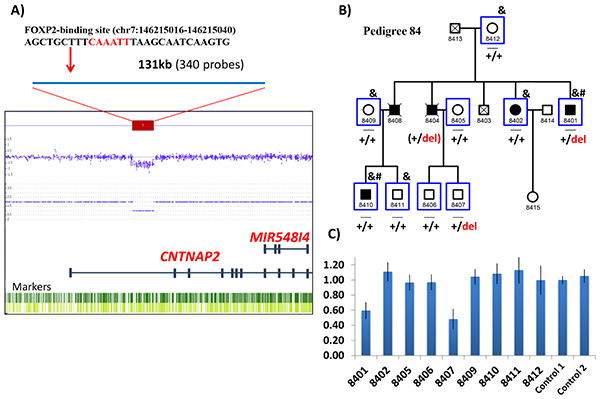
Figure 2. CNV deletion encompassing intron 1 of CNTNAP2 in an extended family with bipolar disorder. A) CytoScan HD array shows the drop in signal intensity of 340 probes, indicating a deletion found in the patient 8401. The position of the FOXP2 binding site within the deletion is shown above. B) The bipolar pedigree includes five patients with bipolar disorder across two generations. Heterozygous deletion carriers are indicated as “+/del” and non-carriers are indicated as “+/+”.C) Gene dosage results of the qPCR experiments validating the deletion in patient 8401, and showing the deletion in unaffected subject 8407.

| Last name | Name | Laboratory | Ext.* | Professional category | |
|---|---|---|---|---|---|
| Calzado González | Ángela | 415.5 | 4731 | Estudiante TFM | |
| Cendón Durán | Andrea | 415.5 | 4731 | Estudiante | |
| García Ortiz | Inés | 415.5.1 | 4731 | ines.garcia(at)cbm.csic.es | Ayudante Investigación |
| Martínez Jiménez | Miriam | 415.5 | 4731 | miriam.martinez(at)cbm.csic.es | Titulado Sup. Actividades Tecn. y Prof.GP1 |
| Rodríguez-Osorio Puigdueta | Cristina | 415 | 4731 | Estudiante TFM | |
| Ruggiero | Alessia | 415.5 | 4731 | Becario Erasmus | |
| Sánchez Moreno | Blanca | 415.5 | 4731 | Estudiante | |
| Somavilla Cabrero | Raúl | 415.5 | 4731 | rsomavilla(at)cbm.csic.es | M1 |
| Toma | Claudio | 415.5.1 | 4731 | claudio.toma(at)cbm.csic.es | Investigador |
| Villacorta Chiotti | Silvana | 207 | 4731 | M2 50% |
Relevant publications:
- Mullins N, et al. (2021) Genome-wide association study of more than 40,000 bipolar disorder cases provides new insights into the underlying biology. Nature Genetics. doi:10.1038/s41588-021-00857-4.
- Toma Claudio*, Shaw AD, Heath A, Pierce KD, Mitchell PB, Schofield PR, Fullerton JM*. (2021) A linkage and exome study of multiplex families with bipolar disorder implicates rare coding variants of ANK3 and additional rare alleles at 10q11-q21. (*, corresponding authors) Journal of Psychiatry and Neuroscience 46(2):E247-E257. doi: 10.1503/jpn.200083.
- Torrico B, Antón-Galindo E, Fernàndez-Castillo N, Rojo E, Ghorbani S, Pineda-Cirera L, Hervás A, Rueda I, Moreno E, Fullerton JM, Casadó V, Buitelaar JK, Rommelse N, Franke B, Reif A, Chiocchetti AG, Freitag C, Kleppe R, Haavik J, Toma Claudio* and Cormand B*. (2020) Involvement of the 14-3-3 gene family in autism spectrum disorder and schizophrenia: genetics, transcriptomics, and functional analyses. Journal of Clinical Medicine. doi.org/10.3390/jcm9061851. *, joint senior and co-corresponding authors.
- Toma Claudio, Shaw AD, Overs BJ, Mitchell PB, Schofield PR, Cooper AA, Fullerton JM. (2020) Coding De novo Variants in Multiplex Bipolar Families are Implicated in Disease Liability. doi: 10.1001/jamanetworkopen.2020.3382 JAMA Network Open; 3(5): e20338216.
- Toma Claudio* (2020) Genetic variation across phenotypic severity of autism. Trends in Genetics 36(4):228-231 doi.org/10.1016/j.tig.2020.01.005 (*, corresponding author)
- Toma Claudio, Díaz-Gay M, Soares de Lima Y, Arnau-Collell C, Franch-Expósito S, Muñoz J, Overs B, Bonjoch L, Carballal S, MD PhD, Ocaña T, Cuatrecasas M, Castells A, Bujanda L, Cubiella J, Balaguer F, Rodríguez-Alcalde D, MD PhD, Fullerton JM, Castellví-Bel S (2019) Identification of a novel candidate gene for serrated polyposis syndrome by performing linkage analysis combined with whole-exome sequencing. Clinical and Translational Gastroenterology
- Toma Claudio, Díaz-Gay M, Franch-Expósito S, Arnau-Collell C, Overs B, Muñoz J, Bonjoch L, Soares de Lima Y, Ocaña T, Cuatrecasas M, Castells A, Bujanda L, Balaguer F, Cubiella J, Caldés T, Fullerton JM, Castellví-Bel S (2020) Using linkage studies combined with whole-exome sequencing to identify novel candidate genes for familial colorectal cancer. International journal of Cancer 15;146(6):1568-1577. doi: 10.1002/ijc.32683
- Torrico B, Shaw AD, Mosca R, Vivo-Luque, Hervas A, Fernàndez-Castillo N, Aloy P, Monica B, Fullerton JM, Cormand B, Toma Claudio*. (2019) Truncating Variant Burden in High Functioning Autism and Pleiotropic Effects of LRP1 Across Psychiatric Phenotypes. doi: 10.1503/jpn.180184 Journal of Psychiatry and Neuroscience 16; 44:1-10. (*, corresponding author)
- Toma Claudio*, Pierce KD, Shaw AD, Heath A, Mitchell PB, Schofield PR, Fullerton JM (2018) Comprehensive cross-disorder analyses of CNTNAP2 suggest it is unlikely to be a primary risk gene for psychiatric disorders. PLoS Genetics Dec 26;14(12):e1007535. doi: 10.1371/journal.pgen.1007535 (*, corresponding author)
- Brandler BM, Antaki D, Gujral M, Kleiber ML, Whitney J, Maile MS, Hong O, Chapman TR, Tan S, Tandon P, Pang T, Tang SC, Vaux KK, Yang Y, Harrington E, Juul S, Turner DJ, Thiruvahindrapuram B, Kaur G, Wang Z, Kingsmore SF, Gleeson JG, Bisson D, Kakaradov B, Telenti A, Venter J C, Corominas R, Toma Claudio, Cormand B, Rueda I, Guijarro S, Messer KS, Nievergelt CM, Arranz MJ, Courchesne E, Pierce K, Muotri AR, Iakoucheva LM, Hervas A, Scherer SW, Corsello C, Sebat J (2018) Paternally inherited cis-regulatory structural variants contribute to autism. Science Apr 20;360(6386):327-331. doi: 10.1126/science.aan2261.
- Toma Claudio, Shaw AD, Allcock RJN, Heath A, Pierce KD, Mitchell PB, Schofield PR, Fullerton JM (2018) An Examination of Multiple Classes of Rare Variants in Extended Families with Bipolar Disorder. Translational Psychiatry. 13;8(1)65.
- Torrico B, Chiocchetti AG, Bacchelli E, Trabetti E, Hervás A, Franke B, Buitelaar JK, Rommelse N, Yousaf A, Duketis E, Freitag CM, Caballero-Andaluz R, Martinez-Mir A, Scholl FG, Ribasés M; ITAN, Battaglia A, Malerba G, Delorme R, Benabou M, Maestrini E, Bourgeron T, Cormand B, Toma Claudio*. (2017) Lack of replication of previous autism spectrum disorder GWAS hits in European populations. Autism Res. 10:202-211. (*, corresponding author)
- Toma Claudio*, Torrico B, Hervás A, Salgado M, Rueda I, Valdés-Mas R, Buitelaar JK, Rommelse N, Franke B, Freitag C, Reif A, Pérez-Jurado LA, Battaglia A, Mazzone L, Bacchelli E, Puente XS, Cormand B. (2015) Common and rare variants of microRNA genes in autism spectrum disorders. World J Biol Psychiatry. 23:1-11 (PMID: 25903372) (*, corresponding author)
- Torrico B, Fernàndez-Castillo N, Hervás A, Milà M, Salgado M, Rueda I, Buitelaar JK, Rommelse N, Oerlemans AM, Bralten J, Freitag CM, Reif A, Battaglia A, Mazzone L, Maestrini E, Cormand B, Toma Claudio*. (2015) Contribution of common and rare variants of the PTCHD1 gene to autism spectrum disorders and intellectual disability. Eur J Hum Genet. 23: 1694-701. (*, corresponding author)
- Toma Claudio, Torrico B, Hervás A, Valdés-Mas R, Tristán A, Maristany M, Padillo V, Romarís P, Arenas C, Puente X, Bayés M, Cormand B (2014) Exome sequencing in multiplex autism families suggests a major role for heterozygous truncating mutations. Mol Psychiatry. 19: 784-90.
Doctoral theses:
- Dr Barbara Torrico Avilés (Universitat de Barcelona, 2012-2015): Molecular genetics of autism: identification of susceptibility variants and functional studies (Co-supervision with Prof. Bru Cormand)




















