Biophysics and systems biology
Research summary:
Our lab focuses on the study of the dynamics of developmental systems combing experimental, theoretical and computational tools. Our main interests are:
- Regulation of stem cell differentiation during developmental processes
The cellular machinery is governed by interacting proteins, genes and metabolites that form complex and highly interconnected networks of interactions. This way, extracellular stimuli triggers pathways of biological events that regulate gene expression, protein activity, and ultimately, cell response. We combine in vivo and in silico approaches to understand how the wiring of the pathways determines the role of the proteins that regulate the differentiation of neurons. We also study how the interplay between mode and rate of division of stem cells is orchestrated. To do that, we use in toto microscopy combined with theoretical tools and algorithms developed in the lab to quantify the effect of key regulators in the balance between proliferation and differentiation during vertebrate neurogenesis. - Nonlinear regulation in pathways and its impact on disease treatment
Small molecule inhibitors display significant potential as treatment for diseases that involve the deregulation of signal transduction pathways. These inhibitors are developed based on their target specificity and binding affinity. We focus on the fact that the numerous signaling proteins and feedback loops in signaling pathways strongly influence the efficiency of pharmacological treatment. The existence of several regulatory positive and negative feedback loops either creates complex dose-responses, desensitization to periodic treatments, or modulation of the drug effect in combinatorial treatments. Our experiments show that the effect of inhibitors strongly depends on the architecture of the targeted pathway, and a detailed characterisation of these nonlinear effects can be useful when designing optimal treatment strategies. - Stochasticity and effect of perturbations in biological networks
Biological networks control cellular behaviour both at the intracellular and at intercellular level. These highly interconnected networks need to perform in the continuously fluctuating and changing cellular microenvironment. Disruption of these networks often leads to aberrant cell behaviour and disease. This leads to some broad questions that we try to address in the lab at different levels: How can these highly nonlinear biological networks integrate and process information? How can they operate robustly in the presence of noise and fluctuations? What are the mechanisms at the network level underlying the adaptation to the different sources of extrinsic and intrinsic noise ? To answer these questions, we use a synthetic biology approach to analyse experimental and computationally the dynamics of fluctuations in minimal networks motifs.
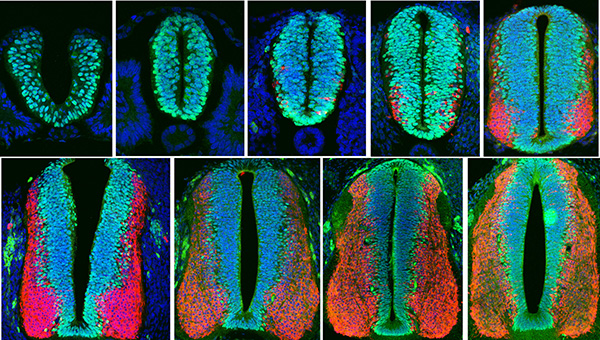
Figure 1: Confocal sections of neural tube of G. Gallus at different developmental times.
Blue: Dapi. Red: Differentiated cells. Green: progenitor cells.

| Last name | Name | Laboratory | Ext.* | Professional category | |
|---|---|---|---|---|---|
| Ballesteros Losa | Ainara | 410 | 4489 | Tco. Sup.Investig. y Laboratorio, GP3 | |
| Ledesma Terrón | Mario | 410 | 4489 | Investigador Doctor | |
| Mazo Durán | Diego | 410 | 4489 | dmazo(at)cbm.csic.es | Contrato Predoctoral |
| Míguez Gómez | David | 410/114.2 | 4489 | dmiguez(at)cbm.csic.es | Profesor Titular Universidad, GA |
| Navarro Martínez | Gemma | 410 | 4489 | Estudiante TFG | |
| Pérez Dones | Diego | 410 | 4489 | Titulado Sup.de Actividades Técn. y Profes. GP1 | |
| Vidakonic | Helena | 410 | 4489 | Estudiante TFM |
Relevant publications:
- M Ledesma-Terrón, N Peralta-Cañadas, DG Míguez, Radial Glial from mammalian developing neocortex can divide symmetrically in vitro, bioRxiv, 707463 (2019)
- D. G. Míguez*, D. Garcia-Morales & F. Casares (2019), Control of size, fate and time by the Hh morphogen in the eyes of flies, Current Topics in Developmental Biology: Volume 137 (in Press), (*shared corresponding authorship)
- D. Garcia-Morales, D. G. Míguez* & F. Casares. Dynamic Hh signaling can generate temporal information during tissue patterning. Development, 146, dev176933 (2019) (*shared corresponding authorship)
- V. L. Doldán Martelli & D.G. Míguez. Drug treatment efficiency depends on the initial state of activation in nonlinear pathways. Scientific Reports, 8, 12495 (2018).
- M. Bernabé-Rubio, G. Andrés, J. Casares-Arias, J. Fernández-Barrera, L. Rangel, N. Reglero-Real, D.C. Gershlick, J.J. Fernández, J. Millán, I. Correas, D.G. Miguez, M.A. Alonso, Novel role for the midbody in primary ciliogenesis by polarized epithelial cells, J. Cell Biology 214, 259–273 (2016)
- A. Valencia-Expósito*, I. Grosheva*, D.G. Míguez*, A. González-Reyes, M. D. Martín-Bermudo, “The Myosin Light Chain Phosphatase is required in epithelial cells to control basal actomyosin oscillations governing tissue elongation”, Nature Communications, 7, 10746 (2016) (*shared 1st authorship)
- D.G. Míguez, “A Branching Process to Characterize the Dynamics of Stem Cell Differentiation”, Scientific Reports, 5, 13265 (2015)
- T. Ruiz-Herrero, J. Estrada, R. Guantes, D. G. Miguez, “A tunable coarse-grained model for ligand-receptor interaction”, PLOS Computational Biology 9, e1003274 (2013).
- D.G. Míguez*, E. Gil-Guiñón, S. Pons and E. Martí, “Smad2 and Smad3 cooperate and antagonize simultaneously in vertebrate neurogenesis”, Journal Of Cell Science 126, 5335 (2013) (*First and corresponding author)
- M. Saade, I. Gutiérrez-Vallejo, G. Le Dréau, M. A. Rabadán, D.G. Miguez, J. Buceta, E. Martí “Sonic hedgehog signaling switches the mode of division in the developing nervous system”, Cell Reports 4, 492 (2013).




















