Regulation of gene expression in Leishmania
Research summary:
Protists of the Leishmania genus include species causing severe diseases in humans, i.e. leishmaniasis. Moreover, Leishmania species are the subject of fundamental interest regarding the evolution of gene regulatory mechanisms, as these organisms have put aside transcriptional regulation. There is currently no acceptable vaccine for preventing leishmaniasis and treatment options are limited. The research activity of our group is focused on studying, on the one hand, genome organization and gene expression in this parasite, and, on the other hand, the immuno-pathological outcomes associated with the infections caused by this parasite. Our aim is contributing knowledge for the development of new methods of controlling this parasitosis.
Massive sequencing techniques have become routine tools in our group for studying molecular mechanisms of gene expression in this parasite. These tasks are being done in close collaboration with Dr Begoña Aguado (head of the Genomics & Massive Sequencing service at CBMSO). Our group has generated new assemblies, improving previous ones, for the genomes of prototypical Leishmania species like L. infantum, causative agent of visceral leishmaniasis in Spain and other countries around the Mediterranean basin, and L. braziliensis, causing mucocutaneous leishmaniasis in South America. For hosting this information, which is actively curated, a web page was created: http://leish-esp.cbm.uam.es. Furthermore, we are using massive RNA sequencing (RNA-seq) for the annotation of transcriptomes of several Leishmania species and to study changes in gene expression. Thus, RNA-seq was used to determine genes involved in parasite resistance against the drugs currently available for treatment of human leishmaniasis. Additionally, we are looking for regulatory cis-elements, often found in the 3’ untranslated regions (UTRs) of mRNAs, associated with RNA-binding proteins (RBPs), as key players in the regulation of gene expression.
Another area of active research, headed by Dr Manuel Soto, is aimed to the study of Leishmania – host immune response interactions, towards the development of prophylactic tools for preventing Leishmania infection. In this regards, a genetically modified-Leishmania mutant line (LiΔHSP70-II) is being analyzed as a promising vaccine; its inoculation induces memory and effector T cell responses against Leishmania able to control subsequent challenges. Finally, as members of the Tropical Diseases network (ISCIII; http://www.ricet.es/es/), our group is engaged in collaborative research activities with different national health institutions.
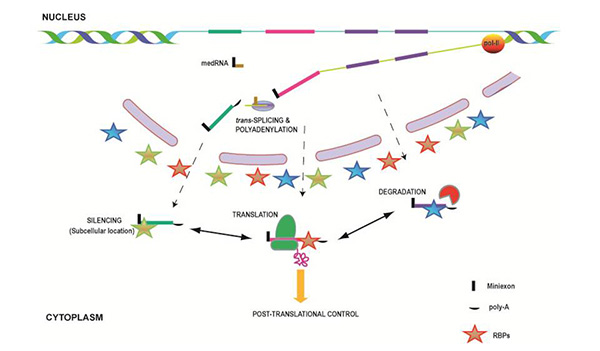
Regulation of gene expression in Leishmania occurs exclusively at the post-transcriptional level.

| Last name | Name | Laboratory | Ext.* | Professional category | |
|---|---|---|---|---|---|
| Adán Jiménez | Javier | 302 | 4647 | javier.adan(at)cbm.csic.es | Tco. de Investigación y Laboratorio |
| Alcain Arranz | Paula | 302 | 4617 | Estudiante TFM | |
| Pacheco Hinojosa | Ronny Francisco | 302 | 4647 | rfpacheco(at)cbm.csic.es | Titulado Sup.Inv.y Laboratorio |
| Requena Rolania | José María | 302 | 4617 | jmrequena(at)cbm.csic.es | Profesor Titular Universidad, GA |
| Sánchez Salvador | Alejandro | 302 | 4647 | alejandro.sanchez(at)cbm.csic.es | Titulado Sup.de Actividades Técn. y Profes. GP1 |
Relevant publications:
- Camacho, E., Gonzalez-de la Fuente, S., Rastrojo, A., Peiro-Pastor, R., Solana, J.C., Tabera, L., Gamarro, F., Carrasco-Ramiro, F., Requena, J.M., and Aguado, B. (2019). Complete assembly of the Leishmania donovani (HU3 strain) genome and transcriptome annotation. Sci Rep 9, 6127. doi: 10.1038/s41598-019-42511-4
- Rastrojo, A., Corvo, L., Lombraña, R., Solana, J.C., Aguado, B., and Requena, J.M. (2019). Analysis by RNA-seq of transcriptomic changes elicited by heat shock in Leishmania major. Sci Rep 9, 6919. doi: 10.1038/s41598-019-43354-9
- Camacho, E., Rastrojo, A., Sanchiz, A., Gonzalez-de la Fuente, S., Aguado, B., and Requena, J.M. (2019). Leishmania Mitochondrial Genomes: Maxicircle Structure and Heterogeneity of Minicircles. Genes (Basel) 10: 758. doi: 10.3390/genes10100758
- Diaz, J.R., Ramirez, C.A., Nocua, P.A., Guzman, F., Requena, J.M., and Puerta, C.J. (2018). Dipeptidyl peptidase 3, a novel protease from Leishmania braziliensis. PLoS One 13, e0190618. doi: 10.1371/journal.pone.0190618
- Rastrojo, A., Garcia-Hernandez, R., Vargas, P., Camacho, E., Corvo, L., Imamura, H., Dujardin, J.C., Castanys, S., Aguado, B., Gamarro, F., and Requena, J.M. (2018). Genomic and transcriptomic alterations in Leishmania donovani lines experimentally resistant to antileishmanial drugs. Int J Parasitol Drugs Drug Resist 8, 246-264. doi: 10.1016/j.ijpddr.2018.04.002
- Garde, E., Ramirez, L., Corvo, L., Solana, J.C., Martin, M.E., Gonzalez, V.M., Gomez-Nieto, C., Barral, A., Barral-Netto, M., Requena, J.M., Iborra, S., and Soto, M. (2018). Analysis of the Antigenic and Prophylactic Properties of the Leishmania Translation Initiation Factors eIF2 and eIF2B in Natural and Experimental Leishmaniasis. Front Cell Infect Microbiol 8, 112. doi: 10.3389/fcimb.2018.00112
- Gonzalez-de la Fuente, S., Camacho, E., Peiro-Pastor, R., Rastrojo, A., Carrasco-Ramiro, F., Aguado, B., and Requena, J.M. (2018). Complete and de novo assembly of the Leishmania braziliensis (M2904) genome. Mem Inst Oswaldo Cruz 114, e180438. doi: 10.1590/0074-02760180438
- Requena, J.M., Rastrojo, A., Garde, E., Lopez, M.C., Thomas, M.C., and Aguado, B. (2017). Genomic cartography and proposal of nomenclature for the repeated, interspersed elements of the Leishmania major SIDER2 family and identification of SIDER2-containing transcripts. Mol Biochem Parasitol 212, 9-15. doi: 10.1016/j.molbiopara.2016.12.009
- Solana, J.C., Ramirez, L., Corvo, L., de Oliveira, C.I., Barral-Netto, M., Requena, J.M., Iborra, S., and Soto, M. (2017). Vaccination with a Leishmania infantum HSP70-II null mutant confers long-term protective immunity against Leishmania major infection in two mice models. PLoS neglected tropical diseases 11, e0005644. doi: 10.1371/journal.pntd.0005644
- Gonzalez-de la Fuente, S., Peiro-Pastor, R., Rastrojo, A., Moreno, J., Carrasco-Ramiro, F., Requena, J.M., and Aguado, B. (2017). Resequencing of the Leishmania infantum (strain JPCM5) genome and de novo assembly into 36 contigs. Sci Rep 7, 18050. doi: 10.1038/s41598-017-18374-y




















