Chromatin, cancer and the ubiquitin system
Research summary:
DNA replication is a central process in cell biology that mediates the copy of cellular DNA to ensure the faithful transmission of genetic information through cell division. Our group is interested in unraveling the mechanisms of control of DNA replication by the protein modifiers ubiquitin and SUMO. We have described that active replication forks present a SUMO rich environment that is maintained by the action of the deubiquitinase USP7 and is essential for DNA replication (Lecona et al., NSMB 2016). We have proposed that the group modification of the replication machinery plays a role during DNA replication (Lecona and Fernández Capetillo, Bioessays 2016) and the inhibition of USP7 leads to the disassembly of the replisome and a premature activation of the mitotic program that results in cell death (Galarreta et al., under revision). At the moment we are working to determine the functions of the group SUMOylation of the replisome and its interplay with the ubiquitination of replication factors.
One of the functions of SUMO and ubiquitin in chromatin is tagging proteins for their extraction by the AAA ATPase VCP through their recognition by specific partners. Our work has shown that VCP, in association with its adaptor FAF1, cooperates with the deubiquitinase USP7 to control the SUMO and ubiquitin equilibrium during DNA replication (Franz, Valledor et al, under revision). Following this work, we are now trying to understand how VCP affects the dynamics of the replication forks and to determine its targets in the replisome.
In addition, we are interested in the response to the challenges that hinder the advance of replication forks, known as replication stress. In the last years the replication stress response (RSR) has been put forward as a promising target in cancer. Initially the RSR constitutes a barrier for transformation but once cancer cells overcome this barrier their accelerated proliferation increases replication stress and makes them highly dependent on the RSR. We are analyzing the functions of the SUMO pathway and VCP in the RSR and its potential applications for cancer treatment. We are using breast cancer cell models with mutations in BRCA1, a central factor in homologous recombination, to explore the therapeutic potential of inhibitors of SUMOylation and VCP, since these tumors have been shown to be sensitive to the inhibition of the RSR.
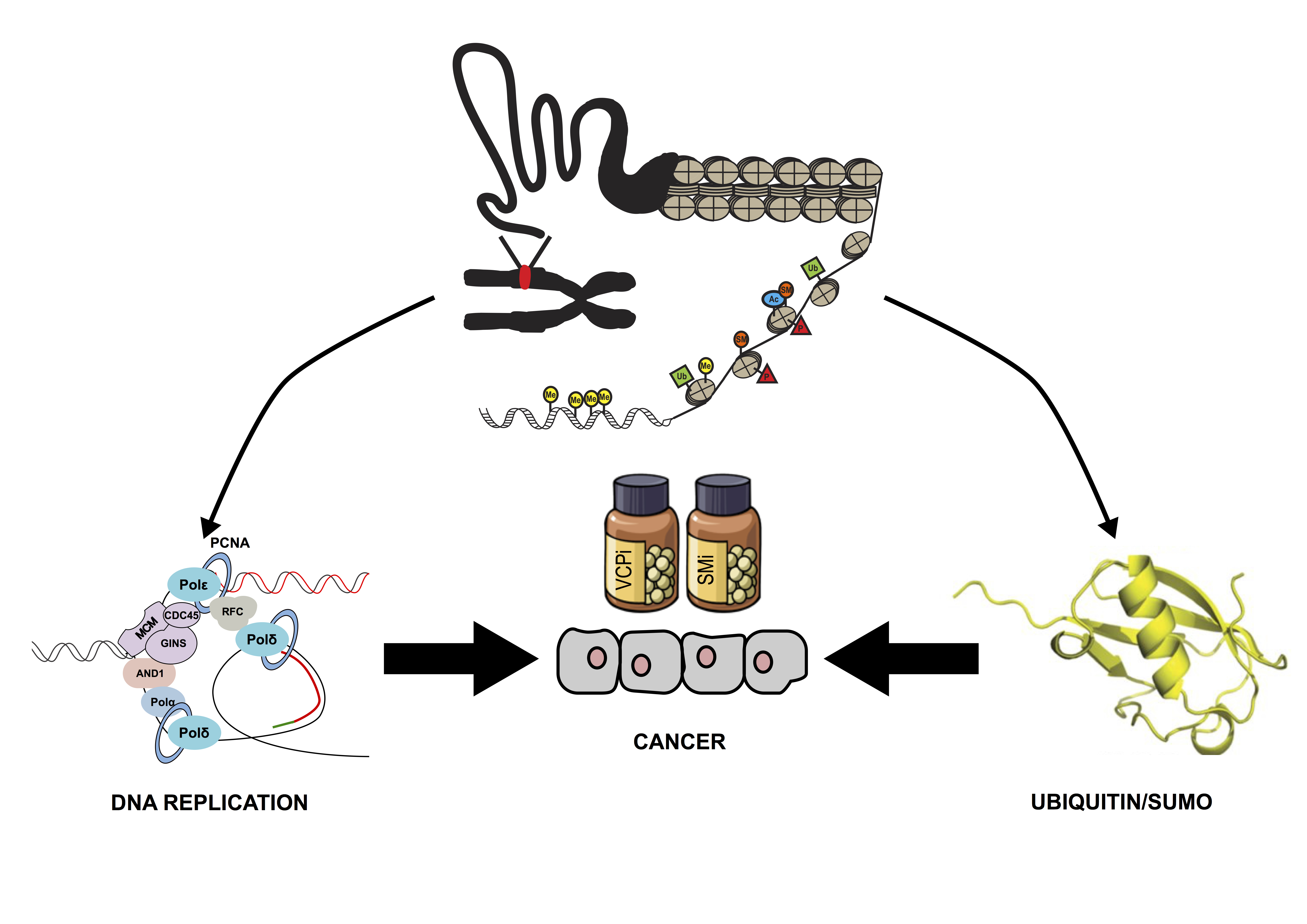
Figure 1. Post-translational modification of proteins in chromatin is essential for the control of DNA replication. Ubiquitin and SUMO are emerging as key modulators of this process and may represent an interesting target for cancer treatment.

| Last name | Name | Laboratory | Ext.* | Professional category | |
|---|---|---|---|---|---|
| Churcher | Scott Brian | 409 | 4486 | sbchurcher(at)cbm.csic.es | M3 Predoc.formación |
| Fuente González | Ana Deodora de la | 409 | 4486 | a.fuente(at)cbm.csic.es | M3 66,66% |
| Gil Plaza | Alba | 409 | 4486 | alba.gil(at)cbm.csic.es | M2 50% |
| Gómez Moya | Alicia | 409 | 4486 | a.gomez.moya(at)cbm.csic.es | M3 Predoc.formación |
| Lecona Sagrado | Emilio | 409 | 4736 | elecona(at)cbm.csic.es | E.Científicos Titulares de Organismos Públicos de Investigación |
| Martín Rufo | Rodrigo | 409 | 4486 | rmartin(at)cbm.csic.es | M2 66,66% |
| Rodríguez Sobrino | Pedro Nicolás | 409 | 4736 | Estudiante. Proyecto Fin de Carrera |
Relevant publications:
- Galarreta A, Valledor P, Ubieto-Capella P, Lafarga V, Zarzuela E, Muñoz J, Malumbres M, Lecona E*, Fernández-Capetillo O. “USP7 limits CDK1 activity throughout the cell cycle” EMBO J (2021) doi: 10.15252/embj.201899692, online ahead of print *Co-corresponding autor
- El Motiam A, de la Cruz-Herrera CF, Vidal S, Seoane R, Baz-Martínez M, Bouzaher YH, Lecona E, Esteban M, Rodríguez MS, Vidal A, Collado M, Rivas C. “SUMOylation modulates the satbility and function of PI·K-p110b” Cell Mol Life Sci (2021) doi: 10.1007/s00018-021-03826-6, online ahead of print.
- Lecona E, Fernández-Capetillo O. “Targeting ATR in Cancer” Nature Reviews Cancer (2018) 18: 586-595
- Mayor-Ruiz C, Olbrich T, Drosten M, Lecona E, Vega-Sendino M, Ortega S, Dominguez O, Barbacid M, Ruiz S, Fernandez-Capetillo O. “ERF deletion rescues RAS deficiency in mouse embryonic stem cells” Genes & Development (2017) 32: 568-576
- Lecona E, Fernández-Capetillo O. “An equilibrium of SUMO and Ubiquitin during DNA replication” Bioessays (2016) 38: 1209-17
- Nieto-Soler M, Morgado-Palacin I, Lafarga V, Lecona E, Murga M, Callen E, Azorin J, Alonso J, López-Contreras AJ, Nussenzweig A, Fernandez-Capetillo O. “Efficacy of ATR inhibitors as single agents in Ewing Sarcoma” Oncotarget (2016) 13: 58759-67.
- Murga M, Lecona E, Kamileri I, Lugli N, Sotiriou S, Días M, Antón ME, Méndez J, Halazonetis T, Fernández-Capetillo O. “POLD3 is haploinsufficient for DNA replication in mice” Molecular Cell (2016) 63: 877-83.
- Lecona E, Rodríguez-Acebes S, Specks J, López-Contreras A, Ruppen I, Murga M, Muñoz J, Méndez J, Fernández-Capetillo O. “USP7 is a SUMO deubiquitinase essential for DNA replication” Nature Structural & Molecular Biology (2016) 23: 270-7 Co-corresponding author
- Lecona E, Narendra V, Reinberg D. “USP7 cooperates with SCML2 to regulate the activity of PRC1” Molecular and Celullar Biology (2015) 35: 1157-68.
- Lecona E, Fernández-Capetillo O. “Replication Stress and Cancer: It Takes Two to Tango” Experimental Cell Research (2014) 329: 26-34.
- Bonasio R*, Lecona E*, Narendra V, Voigt P, Parisi F, Kruger Y, Reinberg D. “Interactions with RNA direct the Polycomb group protein SCML2 to chromatin where it represses target genes” *Equal contribution. eLife (2014) 3:
- Lecona E, Rojas LA, Bonasio R, Johnston A, Fernández-Capetillo O, Reinberg D. “Polycomb protein SCML2 regulates the cell cycle by binding and modulating CDK/CYCLIN/p21 complexes” PLoS Biology (2013) 11: e1001737.
- Bonasio, R., Lecona, E., Reinberg, D. “MBT domains in development and disease” Seminars in Cellular and Developmental Biology (2010) 21:221-230.




















