Gene expression control, patterning and growth during appendage development
Research summary:
The formation of an adult organism from a single fertilized egg is a highly regulated and complex process. It involves the generation of large number of cells with specialized functions that need to be assembled together in tissues and organs. This process, named morphogenesis, requires the specification within developmental fields of territories with the ability to acquire different fates and coordinated cell behaviors. Territorial specification depends on the precise modification of gene expression in both time and space and it is mediated by a specific set of transcription factors encoded by selector and selector-like genes. Selector proteins regulate the formation and identity of tissues and organs activating a special developmental program that is implemented in collective cell behaviors. Cell proliferation, apoptosis or cell shape changes are some of the cellular processes that control the shape and final form an organ.
Functional maintenance of tissues and organs depends on the elimination of defective cells. Misspecified cells or with damage in their DNA are often eliminated by apoptosis. Therefore, a correct balance between cell fate specification, cell proliferation and apoptosis is essential to maintain tissue homeostasis. Defects in the control and coordination of any of these processes are the hallmark of tumor progression.
Our main general aim is to decipher the molecular mechanisms that control cell fate specification and tissue homeostasis during development. To this end we use the development of Drosophila appendages as our model.
We have two objectives in the lab:
1) Specification, patterning and morphogenesis of appendages: We study how legs and wings are specified in the embryo and how they are later subdivided in territories with different cell fates to generate the characteristic adult pattern. We also investigate how these specific gene expression programs are implemented in terms of collective cell behaviors. At the molecular level, a small number of signaling pathways and transcription factors are reiteratively used throughout development to specify and pattern the appendages in Drosophila and in vertebrates.
In this image, a 3D reconstruction of the wing and haltere primordia can be identified (in red) in a developing Drosophila embryo. Epithelial cells are marked in green.
2) Coordination between cell proliferation and apoptosis during tissue homeostasis: The tumor suppressor p53 is a key player in the coordination of cellular responses after multiple stresses such as DNA damage. These responses include the cell cycle arrest, the activation of DNA repair mechanisms and, if the damage is too severe, apoptosis. Defects in the ability to trigger and coordinate any of these responses are a major cause of cancer development. In this work, we use Drosophila to study the connection between cell cycle progression and the apoptotic response after DNA damage.
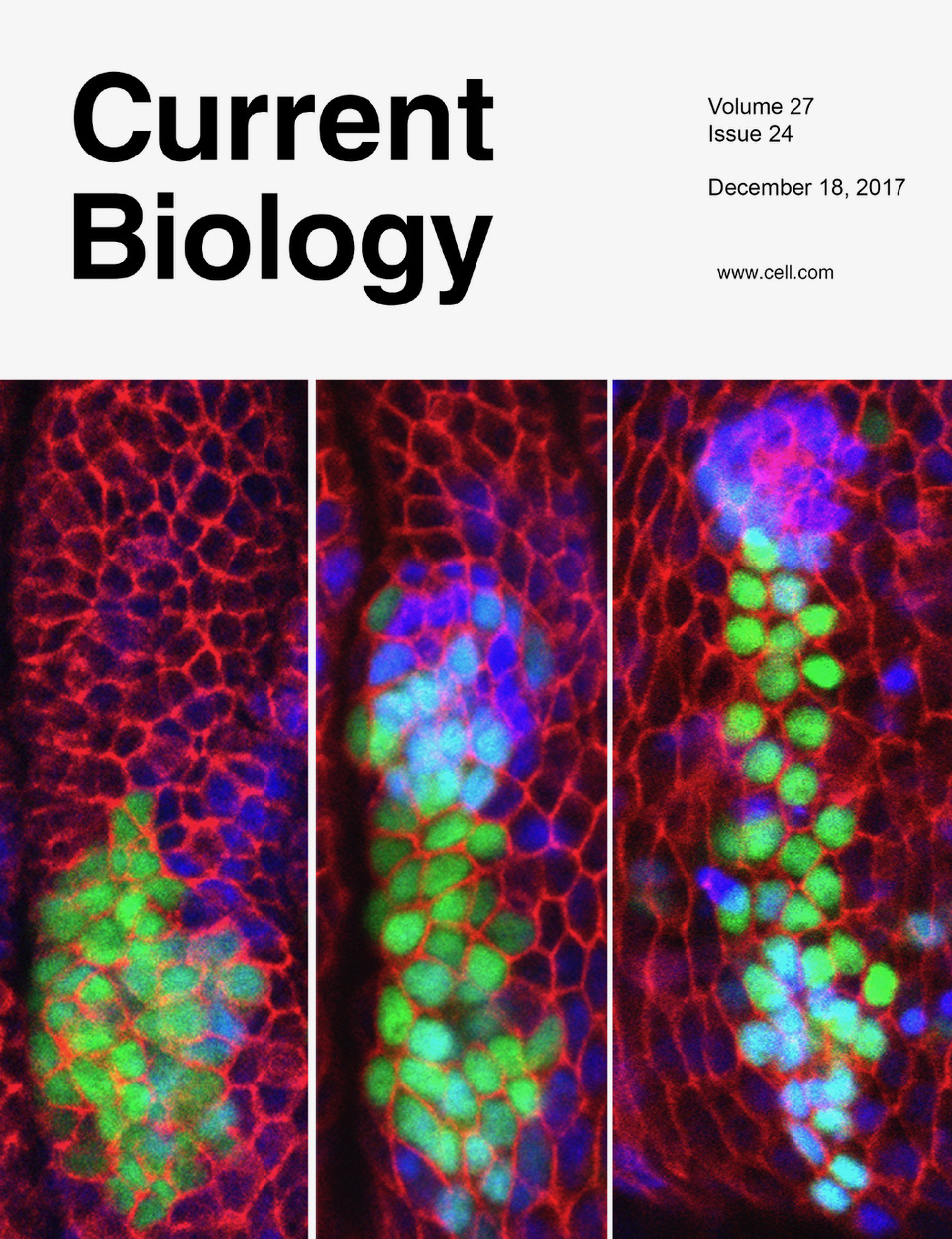
Figure 1: Time course of the specification of the wing and leg primordia in a Drosophila embryo. A group of cells from the leg primordia (green) give rise to part of the wing (blue). Red marks cell membranes.
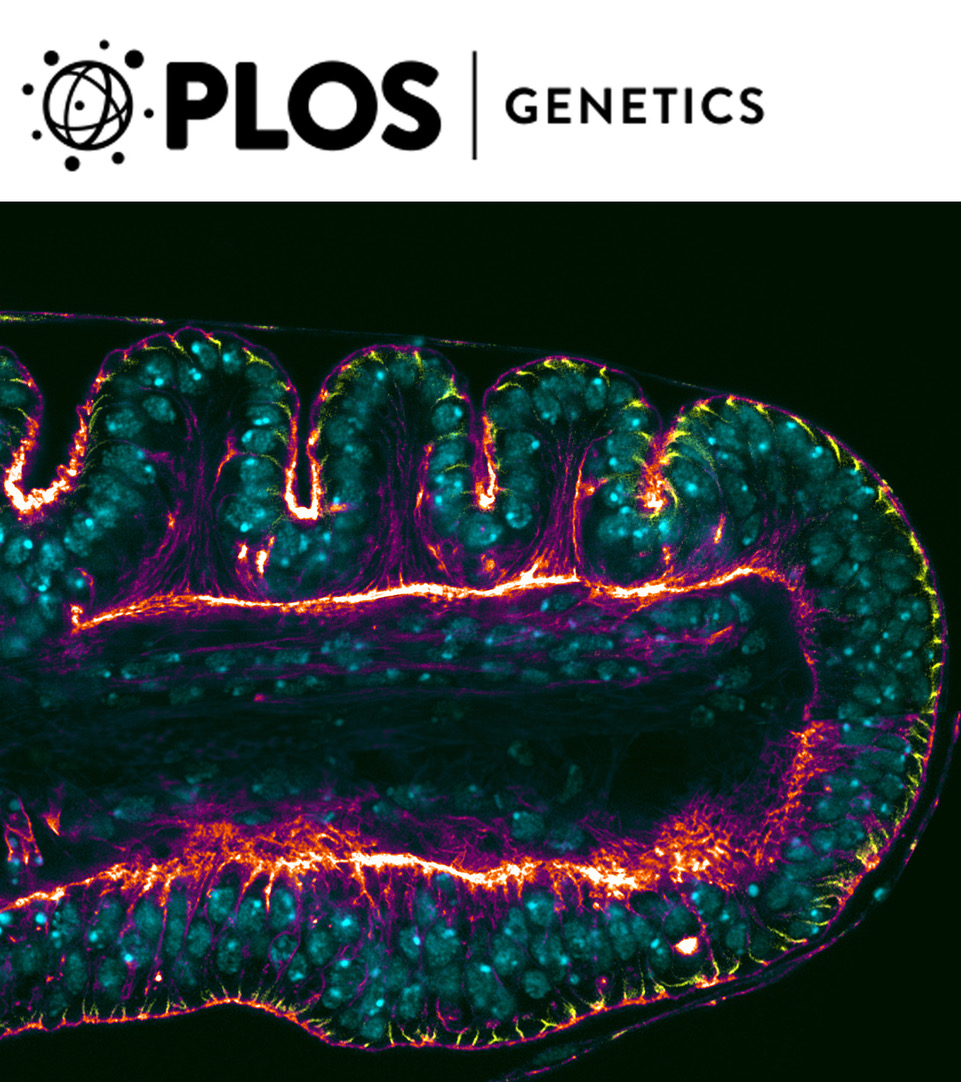
Figure 2: The image is a composition of a wild-type leg imaginal disc showing the folds that prefigure the tarsal joints (upper side) and a dysf mutant leg where the folds are absent (lower side). Dlg (yellow), Nuclear marker (Blue) and F-Actin (Red-White).

Figure 3: Leg phenotypes obtained after a progressive reduction in the dose of Sp family genes in Drosophila.

| Last name | Name | Laboratory | Ext.* | Professional category | |
|---|---|---|---|---|---|
| Castellanos Aguilar | César | 421 | 4436 | cesar.castellanos(at)cbm.csic.es | M3 Predoc.formación |
| Estella Sagrado | Carlos | 424 | 4402 | cestella(at)cbm.csic.es | E.Científicos Titulares de Organismos Públicos de Investigación |
| Kelleher López | Inés | 421 | 4436 | ikelleher(at)cbm.csic.es | M3 Predoc.formación |
Relevant publications:
- 1: Ruiz-Losada M, González R, Peropadre A, Gil-Gálvez A, Tena JJ, Baonza A, Estella C. Coordination between cell proliferation and apoptosis after DNA damage in Drosophila. Cell death and differentiation, 10.1038/s41418-021-00898-6. 25 Nov. 2021, doi:10.1038/s41418-021-00898-6. PMID: 34824391
- 2: Ruiz-Losada M, Pérez-Reyes C, Estella C. Role of the Forkhead Transcription Factors Fd4 and Fd5 During Drosophila Leg Development. Front Cell Dev Biol. 2021 Aug 2;9:723927. doi: 10.3389/fcell.2021.723927. PMID: 34409041; PMCID: PMC8365472.
- 3: Velarde SB, Quevedo A, Estella C, Baonza A. Dpp and Hedgehog promote the glial response to neuronal apoptosis in the developing Drosophila visual system. PLoS Biol. 2021 Aug 11;19(8):e3001367. doi: 10.1371/journal.pbio.3001367. PMID: 34379617; PMCID: PMC8396793.
- 4: Blom-Dahl D, Córdoba S, Gabilondo H, Carr-Baena P, Díaz-Benjumea FJ, Estella C. In vivo analysis of the evolutionary conserved BTD-box domain of Sp1 and Btd during Drosophila development. Dev Biol. 2020 Oct 1;466(1-2):77-89. doi: 10.1016/j.ydbio.2020.07.011. Epub 2020 Jul 29. PMID: 32738261.
- 5: Córdoba S, Estella C. Role of Notch Signaling in Leg Development in Drosophila melanogaster. Adv Exp Med Biol. 2020;1218:103-127. doi:10.1007/978-3-030-34436-8_7. PMID: 32060874.
- 6: Córdoba S, Estella C. The transcription factor Dysfusion promotes fold and joint morphogenesis through regulation of Rho1. PLoS Genet. 2018 Aug 6;14(8):e1007584. doi: 10.1371/journal.pgen.1007584. PMID: 30080872; PMCID: PMC6095628.
- 7: Ruiz-Losada M, Blom-Dahl D, Córdoba S, Estella C. Specification and Patterning of Drosophila Appendages. J Dev Biol. 2018 Jul 14;6(3):17. doi: 10.3390/jdb6030017. PMID: 30011921; PMCID: PMC6162442.
- 8: Requena D, Álvarez JA, Gabilondo H, Loker R, Mann RS, Estella C. Origins and Specification of the Drosophila Wing. Curr Biol. 2017 Dec 18;27(24):3826-3836.e5. doi: 10.1016/j.cub.2017.11.023. Epub 2017 Dec 7. PMID: 29225023; PMCID: PMC5757315.
- 9: Córdoba S, Requena D, Jory A, Saiz A, Estella C. The evolutionarily conserved transcription factor Sp1 controls appendage growth through Notch signaling. Development. 2016 Oct 1;143(19):3623-3631. doi: 10.1242/dev.138735. Epub 2016 Aug 30. PMID: 27578786.
- 10: Córdoba S, Estella C. The bHLH-PAS transcription factor dysfusion regulates tarsal joint formation in response to Notch activity during drosophila leg development. PLoS Genet. 2014 Oct 16;10(10):e1004621. doi: 10.1371/journal.pgen.1004621. PMID: 25329825; PMCID: PMC4199481.
- 11: Bieli D, Kanca O, Requena D, Hamaratoglu F, Gohl D, Schedl P, Affolter M, Slattery M, Müller M, Estella C. Establishment of a Developmental Compartment Requires Interactions between three Synergistic Cis-regulatory Modules. PLoS Genet. 2015 Oct 15;11(10):e1005376. doi: 10.1371/journal.pgen.1005376. PMID: 26468882; PMCID: PMC4607503.
Collaborators:
Richard Mann (Columbia University, NYC)
Markus Affolter (Biozentrum, Basel)
Matthew Slattery (Univ. of Minnesota, MN)
Jose Felix de Celis (CBMSO)
Antonio Baonza (CBMSO)




















